Research News
Stem cell fate choice: determined in an instant
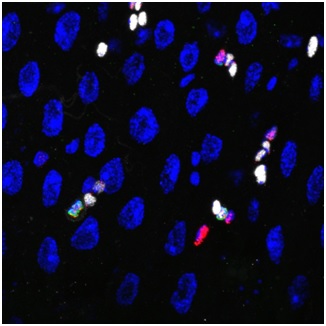
Caption: Stem cells are capable of generating multiple types of cells, but it is unclear how these different types of cells are generated from a common stem cell pool. This study shows that, in Drosophila Intestinal stem cells, a transient activation of a fate inducer determines both the type and number of cells that the stem cells will generate.
January 15, 2017, Dr. Rongwen Xi’s laboratory published online a research article in Nature Cell Biology entitled "Transient Scute activation via a self-stimulatory loop directs enteroendocrine cell pair specification from self-renewing intestinal stem cells". In this article, the authors reported that a transient activation of a fate inducer determines both the type and number of committed progenitor cells from Drosophila intestinal stem cells.
Even as adults, we have stem cells throughout our bodies that are responsible for maintaining many of our tissues. These adult stem cells constantly divide and produce daughter cells, which, through a process called differentiation, become multiple types of mature cells. Scientifically, it’s unclear how these diverse distinct cell types are specified from common stem cell pools. To study this question, the authors investigated cell fate in a multipotent intestinal stem cell (ISC) experimental model from adult fruit flies. The default mode for cell fate is that ISCs differentiate into enterocytes. However, there is a less-well-understood mode in which ISCs differentiate into enteroendocrine cells (EEs). Previous studies have revealed that the four-gene cluster acheate-scute complex (AS-C) act as EE-fate-determination factors. It is also known that one of the AS-C genes—scute (sc)—is both necessary and sufficient for EE specification. Nevertheless, important questions remain about both the molecular and cellular mechanisms through which Sc functions in EE fate decision, and we do not yet know how Sc is regulated to control EE fate.
In their study, the Xi laboratory discovered that EE fate is driven by the transient upregulation of Sc expression in ISCs. Based on the finding that Sc is required for EE generation from ISCs, they temporally knocked down sc starting from the pupal stage, and this process produced flies with midguts lacking EE cells. They used these EE-less midguts to examine the process of EE production by using temperature shift to re-introduce Sc expression in the midgut. With this assay, they discovered that (i) ISCs actually undergo an initial division to generate a new EE progenitor cell which they have termed EEP cell, and (ii) the EEP then undergoes one final round of cell division to produce a pair of EEs.
To further analyze this two-step cell division process, they genetically overexpressed sc in ISCs and monitored the cellular events in a time-course experiment. This transient sc expression caused a rapid cell division response and also induced expression of the EE-marker gene Pros, which is, however, a potent cell-cycle inhibitor. These findings enable the Xi lab to precisely define the regulatory circuitry that directs the formation of a pair of EEs from each ISC via two stepwise cell divisions: an asymmetric division of ISC to generate an EEP, followed by EEP division to produce the EE pair.
Next, to visualize the expression of Sc in midgut, they generated a green fluorescent protein (GFP) tagged line for Sc with the powerful CRISPR-Cas9 genome editing technique. The Sc-GFP fusion protein is weakly expressed in most ISCs, but is expressed at higher levels in ~15% of the ISCs. Follow-up cell lineage tracing studies with an identified Sc-Gal4 line revealed that the immediate daughter cells of Sc-GAL4+ ISCs were mainly EEs; however, these ISCs re-assume their default EC-producing fate as Sc expression is rapidly downregulated.
Mechanistically, it is intriguing how such transient upregulation of Sc in ISCs occurs. By combining genetic assays, RNA-seq analysis, ChIP-seq, and targeted DamID analysis, the Xi group discovered that two feedback regulatory loops control the transient upregulation of Sc in ISCs prior to EE fate commitment. There is a transcriptional self-stimulation loop that allows Sc to gradually build up and eventually reach a high level to induce EEP specification and there is a negative feedback regulation loop between Sc and enhancer of split complex (E(spl)) genes that returns sc expression back to the baseline level.
Given that negative feedback is a common mechanism underlying biochemical oscillations in virtually all organisms, these feedback loops could plausibly be the driver of an oscillatory expression pattern for Sc in ISCs; in theory such oscillatory expression could potentially serve as an internal timer for periodic production of EEs from ISCs. This would be similar to what is known about the circadian clocks, a biological research field that was recently honored with the 2017 Nobel Price for Physiology or Medicine. Future cellular and molecular analysis, likely in combination with in vivo live imaging work will allow further testing and refining of the oscillation model proposed in their study, and such experiments will determined whether and how any internal timer is regulated by certain endogenous and/or environmental cues.
Jun Chen from the Xi laboratory is the first author of the study. Other contributors include Dr. Na Xu, Dr. Chenhui Wang, Dr. Pin Huang, and PhD student Zhen Jin from the Xi laboratory, Huanwei Huang and Dr. Tao Cai from the NIBS sequencing center, as well as Dr. Zhongsheng Yu and Dr. Renjie Jiao from the Institute of Biophysics of the Chinese Academy of Sciences. Dr. Rongwen Xi is the corresponding author. The research was supported by grants from the Chinese Ministry of Science and Technology, National Natural Science Foundation, and the municipal government of Beijing, and was conducted at the National Institute of Biological Sciences, Beijing.



